The workshop focuses on the publication, curation, retrieval, and usage of kinetic data from the reaction kinetics database SABIO-RK and on the use of data in modeling. There will be experience reports from scientists who successfully used experimental data to formulate or verify biological hypotheses with the computer, and you will experience how experimental data can be used with computational models.
Programme: de.NBI Systems Biology Service Center (de.NBI-SysBio)
SEEK ID: https://fairdomhub.org/projects/49
Public web page: http://www.h-its.org/event/kinetics-on-the-move/
Organisms: Bacillus subtilis, Acidithiobacillus caldus, Arabidopsis thaliana
FAIRDOM PALs: No PALs for this Project
Project created: 19th May 2016
Related items
- People (40)
- Programmes (1)
- Institutions (8)
- Investigations (2+1)
- Studies (2+1)
- Assays (3+1)
- Data files (4+6)
- Models (1+3)
- SOPs (1)
- Publications (7)
- Presentations (4+3)
- Events (2+1)
Projects: Kinetics on the move - Workshop 2016
Institutions: Kinetics on the move Workshop at HITS
Projects: SysMetEx, Kinetics on the move - Workshop 2016
Institutions: Università della Svizzera Italiana
Expertise: ODE modelling of biological interaction network, Bioinformatics
Tools: Python, c++, Java, bash, standard bioinformatic tools
Projects: FAIRDOM, Early Metabolic Injury (LiSyM-EMI - Pillar I), Chronic Liver Disease Progression (LiSyM-DP - Pillar II), Regeneration and Repair in Acute-on-Chronic Liver Failure (LiSyM-ACLF - Pillar III), LiSyM Core Infrastructure and Management (LiSyM-PD), Liver Function Diagnostics (LiSyM-LiFuDi - Pillar IV), Model Guided Pharmacotherapy In Chronic Liver Disease (LiSyM-MGP), Multi-Scale Models for Personalized Liver Function Tests (LiSyM-MM-PLF), The Hedgehog Signalling Pathway (LiSyM-JGMMS), Molecular Steatosis - Imaging & Modeling (LiSyM-MSIM), Kinetics on the move - Workshop 2016, Example use cases, FAIRDOM user meeting, MS_DILI, COMBINE Multicellular Modelling, FAIRDOM & LiSyM & de.NBI Data Structuring Training, EnzymeML, GMDS Project Group "FAIRe Dateninfrastrukturen für die Biomedizinische Informatik", FAIRDOM Community Workers, COVID-19 Disease Map, COVID-19 related studies and tools in Germany, nfdi4health - German National Research Data Infrastructure for Personal Health Data, ModeleXchange initiative, SDBV/HITS
Institutions: Heidelberg Institute for Theoretical Studies (HITS gGmbH)
 https://orcid.org/0000-0002-8683-7084
https://orcid.org/0000-0002-8683-7084
Data management and standardization expert for systems biology and systems medicine, responsible for the data management user requirements and user contacts within the German LiSyM network (Liver Systems Medicine: http://lisym.org/) and associated to the FAIRDOM team. Involved in different standardization initiatives and committees, i.e. COMBINE (http://co.mbine.org), ISO/TC 276 Biotechnology (https://www.iso.org/committee/4514241.html), European COST action CHARME (http://www.cost-charme.eu) and ...
Projects: Kinetics on the move - Workshop 2016
Institutions: Novosibirsk State University
 https://orcid.org/0000-0001-8957-6142
https://orcid.org/0000-0001-8957-6142
Projects: Kinetics on the move - Workshop 2016
Institutions: Kinetics on the move Workshop at HITS
Projects: de.NBI-SysBio, Kinetics on the move - Workshop 2016, Example use cases
Institutions: Heidelberg Institute for Theoretical Studies (HITS gGmbH), University of Rostock
Expertise: Data Management, Databases
Tools: MySQL, Java, graphdatabase, Neo4J
Projects: Kinetics on the move - Workshop 2016
Institutions: Kinetics on the move Workshop at HITS
Expertise: chemoinformatics, drug design, molecular modeling, Scientific Computing
Projects: SysMO-LAB, de.NBI-SysBio, Kinetics on the move - Workshop 2016, Example use cases
Institutions: Heidelberg Institute for Theoretical Studies (HITS gGmbH)
Expertise: Biochemistry, Biology
Tools: Databases, Data Management
I'm a biologist working in the field of scientific datases as a biocurator.
Projects: Kinetics on the move - Workshop 2016
Institutions: Kinetics on the move Workshop at HITS
Projects: Kinetics on the move - Workshop 2016, Multi-Scale Models for Personalized Liver Function Tests (LiSyM-MM-PLF), FAIRDOM user meeting, COMBINE Multicellular Modelling, FAIRDOM & LiSyM & de.NBI Data Structuring Training
Institutions: Charité University Medicine Berlin, Humboldt-Universität zu Berlin, Humboldt University Berlin
 https://orcid.org/0000-0003-1725-179X
https://orcid.org/0000-0003-1725-179X
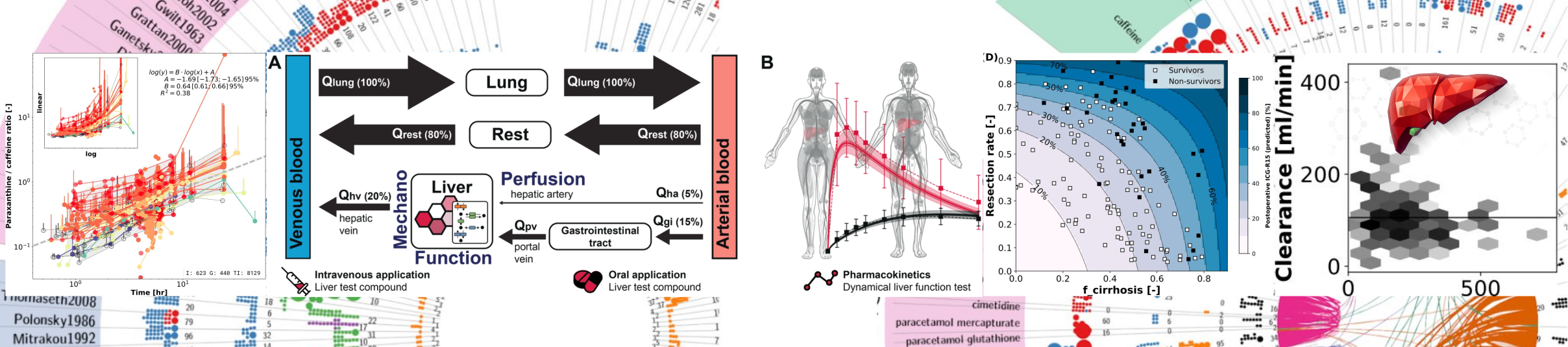
We are investigating liver metabolism and function with the help of computational models and methods.
Group Leader Dr. Matthias König
Institute for Theoretical Biology Humboldt-University Berlin Philippstraße 13, 10115 Berlin, Germany phone +49 30 2093-98435 koenigmx@hu-berlin.de https://www.livermetabolism.com
The König group works on computational modeling, data science, data management, bioinformatics methods and machine learning on ...
Projects: SysMO DB, FAIRDOM, ICYSB 2015 - International Practical Course in Systems Biology, ZucAt, SysMO-LAB, Kinetics on the move - Workshop 2016, Example use cases, FAIRDOM user meeting, ErasysApp Funders, EraCoBiotech 2 nd call proposal preparation, Service to URV Tarragona, Spain with respect to their Safety Assessment of Endocrine Disrupting Chemicals model (Active NOW), FAIRDOM & LiSyM & de.NBI Data Structuring Training, MESI-STRAT, INCOME, Multiscale modelling of state transitions in the host-microbiome-brain network, BESTER, TRALAMINOL, Sustainable co-production, INDIE - Biotechnological production of sustainable indole, Extremophiles metabolsim, PoLiMeR - Polymers in the Liver: Metabolism and Regulation, OLCIR: Optimization of Lung Cancer Therapy with Ionizing Radiation, NAD COMPARTMENTATION, HOTSOLUTE, Stress granules, FAIRDOM Community Workers, GMDS Project Group "FAIRe Dateninfrastrukturen für die Biomedizinische Informatik", Mechanism based modeling viral disease ( COVID-19 ) dynamics in human population, COVID-19 Disease Map, AquaHealth (ERA-BlueBio), LiSyM Core Infrastructure and Management (LiSyM-PD), Early Metabolic Injury (LiSyM-EMI - Pillar I), Regeneration and Repair in Acute-on-Chronic Liver Failure (LiSyM-ACLF - Pillar III), Chronic Liver Disease Progression (LiSyM-DP - Pillar II), Liver Function Diagnostics (LiSyM-LiFuDi - Pillar IV), The Hedgehog Signalling Pathway (LiSyM-JGMMS), Multi-Scale Models for Personalized Liver Function Tests (LiSyM-MM-PLF), Model Guided Pharmacotherapy In Chronic Liver Disease (LiSyM-MGP), Molecular Steatosis - Imaging & Modeling (LiSyM-MSIM), Modelling COVID-19 epidemics, SNAPPER: Synergistic Neurotoxicology APP for Environmental Regulation, SCyCode The Autotrophy-Heterotrophy Switch in Cyanobacteria: Coherent Decision-Making at Multiple Regulatory Layers, SASKit: Senescence-Associated Systems diagnostics Kit for cancer and stroke, CC-TOP, BioCreative VII, MESI-STRAT Review, SDBV/HITS, MESI-Review 2024
Institutions: Heidelberg Institute for Theoretical Studies (HITS gGmbH), FAIRDOM User meeting, Norwegian University of Science and Technology, University of Rostock, University of Innsbruck
 https://orcid.org/0000-0003-3540-0402
https://orcid.org/0000-0003-3540-0402
Expertise: Genetics, Molecular Biology, Bioinformatics, Data Management, Transcriptomics, semantics, Curation, Ontology, Data Modelling
Tools: Cell and tissue culture, Databases, Chip-chip, BioMart, Protege, RightField, SEEK
I am a researcher at the Scientific Databases and Visualization Group at Heidelberg Institute for Theoretical Studies (HITS) , one of the developers of SabioRK - System for the Analysis of Biochemical Pathways - Reaction Kinetics (http://sabiork.h-its.org/) . I am working on design and maintenance of the information systems to store, query and analyse systems biology data; definition and implementation of methods for the integration of data from multiple sources. In **[SySMO-DB ...
Projects: SysMO-LAB, Kinetics on the move - Workshop 2016, de.NBI-SysBio
Institutions: University of Heidelberg
Ursula Kummer is heading the dept. "Modeling of Biological Processes" at the University of Heidelberg.
Projects: Kinetics on the move - Workshop 2016
Institutions: Heidelberg Institute for Theoretical Studies (HITS gGmbH)
Expertise: computational structural biology
Tools: Molecular Dynamics, Data Management, Databases, Data Science, web development, Ruby on Rails, Python
Research associate at HITS, software developer
Projects: Kinetics on the move - Workshop 2016
Institutions: Kinetics on the move Workshop at HITS
Projects: SysMO DB, FAIRDOM, ICYSB 2015 - International Practical Course in Systems Biology, de.NBI-SysBio, Regeneration and Repair in Acute-on-Chronic Liver Failure (LiSyM-ACLF - Pillar III), Early Metabolic Injury (LiSyM-EMI - Pillar I), Chronic Liver Disease Progression (LiSyM-DP - Pillar II), LiSyM Core Infrastructure and Management (LiSyM-PD), Liver Function Diagnostics (LiSyM-LiFuDi - Pillar IV), Molecular Steatosis - Imaging & Modeling (LiSyM-MSIM), Multi-Scale Models for Personalized Liver Function Tests (LiSyM-MM-PLF), Model Guided Pharmacotherapy In Chronic Liver Disease (LiSyM-MGP), The Hedgehog Signalling Pathway (LiSyM-JGMMS), Kinetics on the move - Workshop 2016, Example use cases, SBEpo - Systems Biology of Erythropoietin, FAIRDOM & LiSyM & de.NBI Data Structuring Training, MESI-STRAT, INCOME, EnzymeML, PoLiMeR - Polymers in the Liver: Metabolism and Regulation, MS_DILI, GMDS Project Group "FAIRe Dateninfrastrukturen für die Biomedizinische Informatik", COMBINE Multicellular Modelling, COVID-19 Disease Map, COVID-19 related studies and tools in Germany, nfdi4health - German National Research Data Infrastructure for Personal Health Data, NMTrypI - New Medicines for Trypanosomatidic Infections, ModeleXchange initiative, SNAPPER: Synergistic Neurotoxicology APP for Environmental Regulation, BioCreative VII, SDBV ephemeral data exchanges, The BeeProject, SDBV/HITS, MESI-STRAT Review, MESI-Review 2024
Institutions: Heidelberg Institute for Theoretical Studies (HITS gGmbH)
 https://orcid.org/0000-0002-4980-3512
https://orcid.org/0000-0002-4980-3512
I am group leader of the SDBV (Scientific Databases and Visualisation) group at the HITS gGmbH, the Heidelberg Institute for Theoretical Studies.
I am interested in finding data. Starting with my master's thesis I have always worked on how to store data in a way that you can find it, and how to make sense out of data that has been stored.
Within FAIRDOM I find interesting to help people to store their data in a way that they make sense even after years.
Projects: SysMO DB, FAIRDOM, ICYSB 2015 - International Practical Course in Systems Biology, Kinetics on the move - Workshop 2016
Institutions: Heidelberg Institute for Theoretical Studies (HITS gGmbH)
Expertise: Software Engineering, Data Management, Databases
Tools: Ruby on Rails, MySQL, HTML, Ruby, Java, Javascript
Software developer for FAIRDOM
The German Network for Bioinformatics Infrastructure - de.NBI offers first class bioinformatics services including training and education to users in basic and applied life sciences research. In this network 40 projects belonging to eight service centers provide services that cover a wide variety of methods (genomics, proteomics, ...) and applications (from plants to humans). de.NBI-SysBio is the Systems Biology Service Center of de.NBI. In collaboration with FAIRDOM, de.NBI-SysBio serves the ...
Projects: de.NBI-SysBio, ExtremoPharm, ZucAt, Kinetics on the move - Workshop 2016, Example use cases, MIX-UP, Working Group Nicole Radde, MPIEvolBio-SciComp, SABIO-VIS
Web page: http://www.denbi.de
Research in Systems Biology involves integrating data and knowledge about the dynamic processes in biological systems in order to understand and model them. By connecting fields such as genomics, proteomics, bioinformatics, mathematics, cell biology, genetics, mathematics, engineering and computer sciences, Systems Biology enables discovery of yet unknown principles underlying the functioning of living cells. At the same time, testable and predictive models of complex cellular pathways and ...
Submitter: Ron Henkel
Snapshots: No snapshots
Experimental study for hands on session
Submitter: Olga Krebs
Investigation: Using SEEK and I-S-A structure to integrate ex...
Assays: How to use templates for FAIR experimental data representation, How to use templates to structure experimental data 222
Snapshots: No snapshots
Submitter: Ron Henkel
Investigation: Hands-on: Model Management in SEEK
Snapshots: No snapshots
Submitter: Ron Henkel
Biological problem addressed: Cell Cycle
Investigation: Hands-on: Model Management in SEEK
Organisms: No organisms
Models: BIOMD0000000005 from BioModels Database
SOPs: No SOPs
Data files: SEDML for BIOMD0000000005
Snapshots: No snapshots
Here we will use JERM templates with embedded ontologies to describe and annotate example data
Submitter: Olga Krebs
Assay type: Interactomics
Technology type: Technology Type
Investigation: Using SEEK and I-S-A structure to integrate ex...
Organisms: No organisms
SOPs: Extraction and quantitative analysis of protein...
Data files: Master template, Transcriptomics Template (ArrayExpress Format)
Snapshots: No snapshots
Here we will use JERM templates with embedded ontologies to describe and annotate example data 222
Submitter: Olga Krebs
Assay type: Experimental Assay Type
Technology type: Technology Type
Investigation: Using SEEK and I-S-A structure to integrate ex...
Organisms: Bacillus subtilis : test 1 (insertion xyl / wild-type)
SOPs: Extraction and quantitative analysis of protein...
Data files: 0804_shake-flask-sigB-starvation
Snapshots: No snapshots
Master template for hands on session
Creator: Olga Krebs
Submitter: Olga Krebs
Investigations: Using SEEK and I-S-A structure to integrate ex...
Creator: Ulrike Wittig
Submitter: Ulrike Wittig
Investigations: No Investigations
Studies: No Studies
Assays: No Assays
Creator: Ulrike Wittig
Submitter: Ulrike Wittig
Investigations: No Investigations
Studies: No Studies
Assays: No Assays
Creator: Ulrike Wittig
Submitter: Ulrike Wittig
Investigations: No Investigations
Studies: No Studies
Assays: No Assays
This is BIOMD0000000005.
Creators: Ron Henkel, Dagmar Waltemath
Submitter: Ron Henkel
Model type: Ordinary differential equations (ODE)
Model format: SBML
Environment: Copasi
Organism: Not specified
Investigations: Hands-on: Model Management in SEEK
A Standard Operating Procedure (SOP) is a document consisting of step-by-step information on how to execute a task. An existing SOP may need to just be modified and updated.
Creator: Olga Krebs
Submitter: Deleted submitter
Abstract
Authors: Wolfgang Müller, Meik Bittkowski, Martin Golebiewski, Renate Kania, Maja Rey, Andreas Weidemann, Ulrike Wittig
Date Published: 1st Mar 2017
Publication Type: Journal
DOI: 10.1007/s13222-016-0243-4
Citation: Datenbank Spektrum 17(1):21-28
Abstract
Authors: Antoine Buetti-Dinh, Olga Dethlefsen, Ran Friedman, Mark Dopson
Date Published: 26th May 2016
Publication Type: Not specified
DOI: 10.1099/mic.0.000314
Citation: Transcriptomic analysis reveals how a lack of potassium ions increases Sulfolobus acidocaldarius sensitivity to pH changes
Abstract (Expand)
Authors: Y. Hirose, H. Kaida, A. Kawahara, S. Matono, T. Tanaka, S. Kurata, M. Kage, M. Ishibashi, T. Abe
Date Published: 25th May 2016
Publication Type: Not specified
PubMed ID: 27218430
Citation: Nucl Med Commun. 2016 May 23.
Abstract (Expand)
Author: Matthias König
Date Published: 2016
Publication Type: Not specified
DOI: 10.12688/f1000research.9211.1
Citation: F1000Res 5 : 1736
Abstract (Expand)
Authors: M. I. Stefan, S. J. Edelstein, N. Le Novere
Date Published: 31st Jul 2008
Publication Type: Not specified
PubMed ID: 18669651
Citation: Proc Natl Acad Sci U S A. 2008 Aug 5;105(31):10768-73. doi: 10.1073/pnas.0804672105. Epub 2008 Jul 31.
Abstract (Expand)
Author: J. J. Tyson
Date Published: 15th Aug 1991
Publication Type: Not specified
PubMed ID: 1831270
Citation: Proc Natl Acad Sci U S A. 1991 Aug 15;88(16):7328-32.
Abstract
Authors: Thomas H. Crouch, Claude B. Klee
Date Published: 1st Aug 1980
Publication Type: Not specified
DOI: 10.1021/bi00557a009
Citation: Biochemistry 19(16) : 3692
Presentation at the 10th anniversary of SABIO-RK, Heidelberg 2016
Creator: Dagmar Waltemath
Submitter: Dagmar Waltemath
The 16th conference on Computational Methods in Systems Biology (CMSB 2018) will take place on the 12th to 14th September 2018 in Brno, Czech Republic. Its aim is to bring together researchers from across biological, mathematical, computational, and physical sciences who are interested in the study, modelling, simulation, advanced analysis, and design of biological systems.
CMSB 2018 will be hosted at Faculty of Informatics, Masaryk University, Brno. Brno is the city where the modern genetics has ...
Country:  Czechia
Czechia
City: Brno
Here we collect all the presentations, hands on instructions and documentation for Kinetics on the move Workshop on Data for Computational Modeling
Start Date: 30th May 2016
End Date: 31st May 2016
Event Website: http://www.h-its.org/de/research/sdbv/veranstaltungen/kinetics-on-the-move-practical-workshop-on-data-for-computational-modelling/
Country: Not specified
City: Heidelberg
 Overview
Overview Tags
Tags


 Download
Download External Link
External Link