Related items
Projects: IMOMESIC
Institutions: Charité University Medicine Berlin
Projects: MESI-STRAT
Institutions: Charité University Medicine Berlin
Projects: IMOMESIC
Institutions: Charité University Medicine Berlin
Projects: Kinetics on the move - Workshop 2016, Multi-Scale Models for Personalized Liver Function Tests (LiSyM-MM-PLF), FAIRDOM user meeting, COMBINE Multicellular Modelling, FAIRDOM & LiSyM & de.NBI Data Structuring Training
Institutions: Charité University Medicine Berlin, Humboldt-Universität zu Berlin, Humboldt University Berlin
 https://orcid.org/0000-0003-1725-179X
https://orcid.org/0000-0003-1725-179X
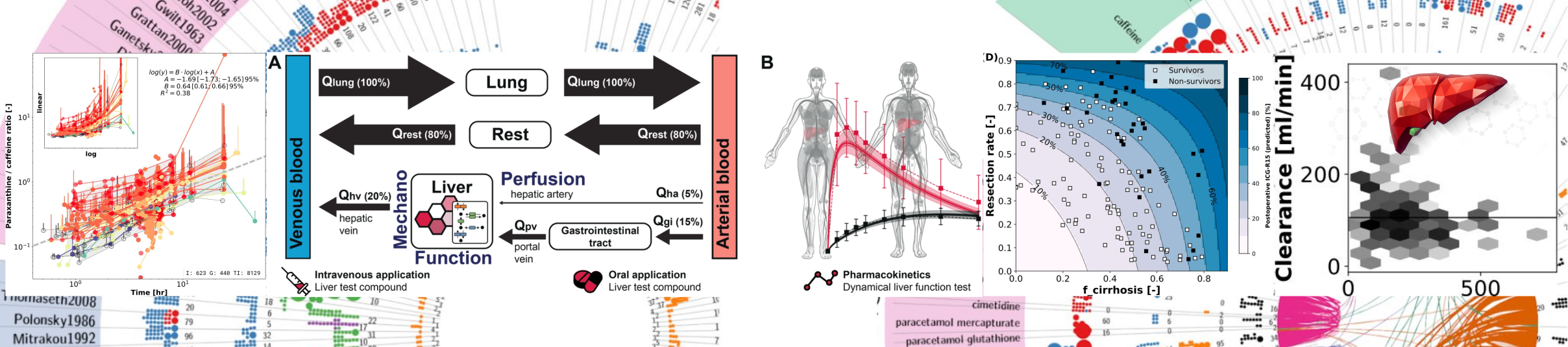
We are investigating liver metabolism and function with the help of computational models and methods.
Group Leader Dr. Matthias König
Institute for Theoretical Biology Humboldt-University Berlin Philippstraße 13, 10115 Berlin, Germany phone +49 30 2093-98435 koenigmx@hu-berlin.de https://www.livermetabolism.com
The König group works on computational modeling, data science, data management, bioinformatics methods and machine learning on ...
Projects: MESI-STRAT
Institutions: Charité University Medicine Berlin
 https://orcid.org/0000-0001-5440-5503
https://orcid.org/0000-0001-5440-5503
Tools: R, Python, Data Science, Bioinformatics, Single Cell analysis
PhD student
Projects: MESI-STRAT
Institutions: Charité University Medicine Berlin
Projects: MESI-STRAT
Institutions: Charité University Medicine Berlin
Projects: IMOMESIC
Institutions: Charité University Medicine Berlin
Projects: MESI-STRAT
Institutions: Charité University Medicine Berlin
The main objective of the ERANET proposal Systems Biology Applications - ERASysAPP (app = application = translational systems biology) is to promote multidimensional and complementary European systems biology projects, programmes and research initiatives on a number of selected research topics. Inter alia, ERASysAPP will initiate, execute and monitor a number of joint transnational calls on systems biology research projects with a particular focus on applications - or in other words so called ...
Projects: SysVirDrug, SysMilk, SysMetEx, MetApp, IMOMESIC, WineSys, CropClock, SYSTERACT, XyloCut, RootBook, ROBUSTYEAST, LEANPROT, ErasysApp Funders
Web page: https://www.cobiotech.eu/about-cobiotech/erasysapp
The DrugLogics initiative comprises a number of different subprojects, all geared towards the development of personalised medicine, or precision medicine
Projects: CausalDB, Colosys, Crossover Research 2.0, NTNU Health Druglogics
Web page: http://druglogics.eu/
LiSyM (Liver Systems Medicine) represents a research network of German centers and institutions, brought together by a 20 Million Euro funding program of the German Government, in which mathematicians, modelers, pharmacologists, molecular biologists and clinical scientists work together to develop a Systems Medicine approach to study early and advanced liver disease. The aim of this unique research program is to acquire and use new experimental data and data from existing data bases to build ...
Projects: Early Metabolic Injury (LiSyM-EMI - Pillar I), Chronic Liver Disease Progression (LiSyM-DP - Pillar II), Regeneration and Repair in Acute-on-Chronic Liver Failure (LiSyM-ACLF - Pillar III), LiSyM Core Infrastructure and Management (LiSyM-PD), Liver Function Diagnostics (LiSyM-LiFuDi - Pillar IV), Model Guided Pharmacotherapy In Chronic Liver Disease (LiSyM-MGP), Multi-Scale Models for Personalized Liver Function Tests (LiSyM-MM-PLF), The Hedgehog Signalling Pathway (LiSyM-JGMMS), Molecular Steatosis - Imaging & Modeling (LiSyM-MSIM), FAIRDOM & LiSyM & de.NBI Data Structuring Training, New LiSyM project
Web page: http://www.lisym.org
The German Network for Bioinformatics Infrastructure - de.NBI offers first class bioinformatics services including training and education to users in basic and applied life sciences research. In this network 40 projects belonging to eight service centers provide services that cover a wide variety of methods (genomics, proteomics, ...) and applications (from plants to humans). de.NBI-SysBio is the Systems Biology Service Center of de.NBI. In collaboration with FAIRDOM, de.NBI-SysBio serves the ...
Projects: de.NBI-SysBio, ExtremoPharm, ZucAt, Kinetics on the move - Workshop 2016, Example use cases, MIX-UP, Working Group Nicole Radde, MPIEvolBio-SciComp, SABIO-VIS
Web page: http://www.denbi.de
MESI-STRAT: Systems Medicine of Metabolic-Signaling Networks -A New Concept for Breast Cancer Patient Stratification. Breast cancer is a complex disease with high prevalence in the European Union and world-wide. 75%-80 of the patients have estrogen receptor-positive (ER)-positive tumors and are treated with endocrine therapies. Endocrine therapies, which block ER-driven tumor growth, show high efficacy. Yet, a significant proportion of the patients will eventually relapse with metastatic breast ...
Programme: This Project is not associated with a Programme
Public web page: http://www.mesi-strat.eu
Organisms: Homo sapiens, Mus musculus, Rattus norvegicus
IMOMESIC - Integrating Modelling of Metabolism and Signalling towards an Application in Liver Cancer
One of the most challenging questions in cancer research is currently the interconnection of metabolism and signalling. An understanding of mechanisms that facilitate the physiological shift towards a proliferative metabolism in cancer cells is considered a major upcoming topic in oncology and is a key activity for future drug development. Due to the complexity of interrelations, a systems biology ...
Programme: ERASysAPP
Public web page: Not specified
Organisms: Homo sapiens
We will contribute to the LiSyM Research Network an open source, freely available and reproducible multiscale model of the human liver from single cell metabolism to whole liver function. The model will be available in existing standards of systems biology, provide standardized interfaces for data integration and be fully annotated to available biological, medical and computational ontologies. All data, models and source code will be shared within the LiSyM Research Network and made available to ...
Programme: LiSyM - Systems Medicine of the Liver
Public web page: https://livermetabolism.com
Organisms: Homo sapiens, Mus musculus, Rattus norvegicus
The workshop focuses on the publication, curation, retrieval, and usage of kinetic data from the reaction kinetics database SABIO-RK and on the use of data in modeling. There will be experience reports from scientists who successfully used experimental data to formulate or verify biological hypotheses with the computer, and you will experience how experimental data can be used with computational models.
Programme: de.NBI Systems Biology Service Center (de.NBI-SysBio)
Public web page: http://www.h-its.org/event/kinetics-on-the-move/
The COLOSYS project aims to develop a deeper understanding of colon cancer networks and convert them into computer models with which it will be better to predict response to treatment. The combination of computational, experimental and clinical testing will provide understanding of drug resistance mechanisms, and allow personalised treatment of colon cancer.
Programme: Druglogics
Public web page: http://www.colosys.org/
Organisms: Homo sapiens
tba.
Programme: Druglogics
Public web page: Not specified
Organisms: Homo sapiens
