Lutz Brusch is heading the research group "Spatio-temporal pattern formation in cells and tissues" at the Centre for Information Services and High Performance Computing of TU Dresden, Germany. The group is co-developing the multi-cellular modelling and simulation framework Morpheus (https://morpheus.gitlab.io) and is collaborating with experimental labs on questions of tissue morphogenesis and regeneration.
Biochemist currently keeping busy as: Research Data Manager (Vrije Universiteit Amsterdam), Software Engineer (Heidelberg University) and member of the SBML Development Team (Caltech).
Projects: Kinetics on the move - Workshop 2016, Multi-Scale Models for Personalized Liver Function Tests (LiSyM-MM-PLF), FAIRDOM user meeting, COMBINE Multicellular Modelling, FAIRDOM & LiSyM & de.NBI Data Structuring Training
Institutions: Charité University Medicine Berlin, Humboldt-Universität zu Berlin, Humboldt University Berlin
 https://orcid.org/0000-0003-1725-179X
https://orcid.org/0000-0003-1725-179X
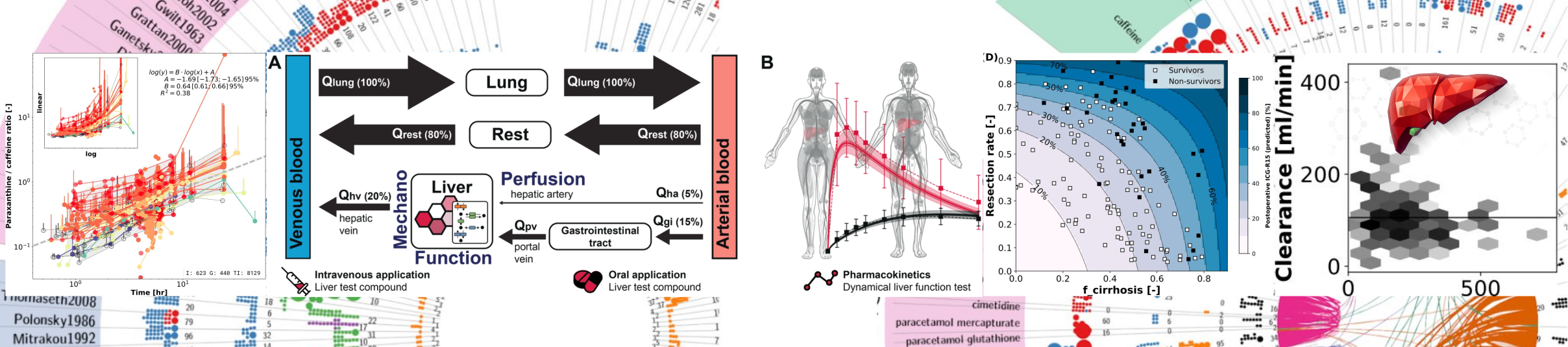
We are investigating liver metabolism and function with the help of computational models and methods.
Group Leader Dr. Matthias König
Institute for Theoretical Biology Humboldt-University Berlin Philippstraße 13, 10115 Berlin, Germany phone +49 30 2093-98435 koenigmx@hu-berlin.de https://www.livermetabolism.com
The König group works on computational modeling, data science, data management, bioinformatics methods and machine learning on ...
Projects: de.NBI-SysBio, GenoSysFat, Kinetics on the move - Workshop 2016, Example use cases, COMBINE Multicellular Modelling, GMDS Project Group "FAIRe Dateninfrastrukturen für die Biomedizinische Informatik", COVID-19 related studies and tools in Germany, nfdi4health - German National Research Data Infrastructure for Personal Health Data
Institutions: University of Rostock, University of Greifswald, University Medicine of Greifswald
 https://orcid.org/0000-0002-5886-5563
https://orcid.org/0000-0002-5886-5563
I am a computer scientist by training with a specialisation on database and information systems. Since December 2018 I am professor of Medical Informatics at the University Medicine in Greifswald, Germany, at the Institute of Community Medicine. My lab focuses on research data management in biomedicine, data integration across health care providers, and provenance of clinical research data items within clinical information systems. Furthermore, I am actively involved in COMBINE standardisation ...
Abstract (Expand)
Authors: Chris J. Myers, Gary Bader, Padraig Gleeson, Martin Golebiewski, Michael Hucka, Nicolas Le Novere, David P. Nickerson, Falk Schreiber, Dagmar Waltemath
Date Published: 1st Dec 2017
Publication Type: InProceedings
Citation: 2017 Winter Simulation Conference (WSC),pp.884-895,IEEE
Abstract (Expand)
Authors: Falk Schreiber, Björn Sommer, Gary D. Bader, Padraig Gleeson, Martin Golebiewski, Michael Hucka, Sarah M. Keating, Matthias König, Chris Myers, David Nickerson, Dagmar Waltemath
Date Published: 13th Jul 2019
Publication Type: Journal
Citation: Journal of Integrative Bioinformatics 16(2)
Abstract (Expand)
Authors: Falk Schreiber, Björn Sommer, Tobias Czauderna, Martin Golebiewski, Thomas E. Gorochowski, Michael Hucka, Sarah M. Keating, Matthias König, Chris Myers, David Nickerson, Dagmar Waltemath
Date Published: 29th Jun 2020
Publication Type: Journal
Citation: Journal of Integrative Bioinformatics 0(0)
Abstract (Expand)
Authors: Dagmar Waltemath, Martin Golebiewski, Michael L Blinov, Padraig Gleeson, Henning Hermjakob, Michael Hucka, Esther Thea Inau, Sarah M Keating, Matthias König, Olga Krebs, Rahuman S Malik-Sheriff, David Nickerson, Ernst Oberortner, Herbert M Sauro, Falk Schreiber, Lucian Smith, Melanie I Stefan, Ulrike Wittig, Chris J Myers
Date Published: 29th Jun 2020
Publication Type: Journal
Citation: Journal of Integrative Bioinformatics 0(0)
Abstract (Expand)
Authors: F. Schreiber, G. D. Bader, P. Gleeson, M. Golebiewski, M. Hucka, N. Le Novere, C. Myers, D. Nickerson, B. Sommer, D. Walthemath
Date Published: 12th Feb 2017
Publication Type: Not specified
PubMed ID: 28187405
Citation: J Integr Bioinform. 2016 Dec 18;13(3):289. doi: 10.2390/biecoll-jib-2016-289.
Written and presented by Dagmar Waltemath (University of Rostock) as part of the Reproducible and Citable Data and Models Workshop in Warnemünde, Germany. September 14th - 16th 2015.
Creators: Natalie Stanford, Dagmar Waltemath
Submitter: Natalie Stanford
 External Link
External Link